iPRG Study Results
The Proteome Informatics Research Group (iPRG) 2013 Study evaluated the benefits of using databases derived from RNA-Seq data for peptide and protein identification from tandem mass spectrometry data.
Participating researchers were required to assign the CID spectra present in the sample with < 1% false discovery rate (FDR) at the unique peptide/sequence level for matches to multiple different target databases.
The graph below presents the performance of all 17 submissions.
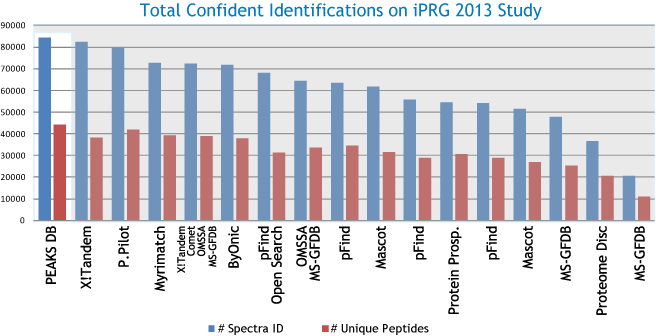
Notice the significant number of spectra and unique peptides identified by PEAKS with high confidence.
In conclusion, the study showed that:
- Confident interpretations were reported for a surprisingly high percentage (82%) of spectra acquired.
- Much higher agreement (and better reliability?) for SNV identifications compared to novel sequence IDs.
- Consensus among results from same participant/lab clearly inflated consensus for novel sequence identification.
- Evidence for high FDR among extra sequence identifications for some participants (decoy database matches concentrated among extra identifications).
- Many SNV and some novel sequence IDs are found in other reference databases.

