
 PEAKS CMD™
PEAKS CMD™
Complete Command Line-Based Solution for Proteomics
Enabling users to batch-process data on a dedicated resource without manual intervention
Compatibility
Exporting
The primary objectives of PEAKS CMD is to provide a smooth workflow with enhanced analytical capabilities. It is also necessary to support downstream analysis, therefore users have the ability to export standard supported data formats such as pepXML and mzIdentML.
Algorithms
Industry Leading de novo Sequencing
PEAKS is the preferred de novo sequencing software for identifying novel peptides and studying unsequenced organisms. Even when the peptide sequence is in a database, de novo sequencing can greatly improve the database search identification.
PEAKS has been reported as the best-performing de novo sequencing software in multiple independent publications.
Sensitive and Accurate Database Searching
Database searching is a standard way to identify the peptides whose sequences are in a database. It is also an important function of the PEAKS software. While database searching cannot discover new peptides, it is a great way to confirm the existence of expected peptides and PTMs. It also serves the foundation of many other proteomics analyses, such as protein quantification and deep replication.
Advanced Post Translational Modification Searching
Post-translational modifications (PTMs) are crucial to the functions of proteins. Their accurate and sensitive characterization is of great importance to every proteomics researcher. Since the PTM information is usually absent or incomplete in today’s protein databases, the identification of PTMs poses a number of challenges to the traditional database search approach. The PEAKS software includes the advanced PEAKS PTM algorithm to meet these challenges and is the only commercial software that can identify unspecified variable PTMs.
Sequence Tag Homology Searching
Biological samples commonly contain proteins with slightly different sequences to those protein databases. This is frequently caused by polymorphism, antibody diversity, database errors, and cross-species database searching. Ignoring those mutated peptides can potentially lead to an oversight of a potential biomarker, an error in antibody confirmation, or simply low coverage of proteins. To handle mutations, PEAKS software includes the SPIDER algorithm that is specially designed to detect peptide mutations and perform cross-species homology search.
Configuration
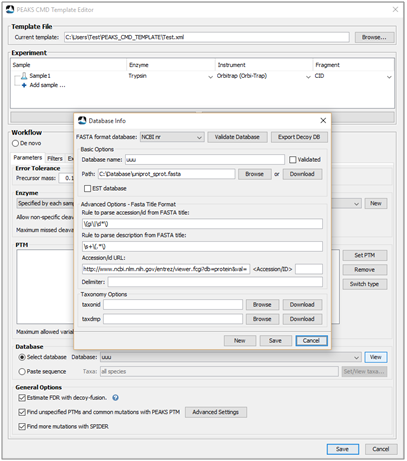
Designed for integration into existing and customized analytical pipelines using standard protocols. The workflow includes configurations of MS file loading, search types, parameters, protein/peptide filtration, and result exporting, is customized by users and stored in a template file. To run PEAKS CMD tasks, users can easily create new (or edit an existing) template files for immediate, recurring or future use.
Licence Information
The PEAKS CMD licence can be scaled to address your lab’s requirements.

