Proteomics is the study of proteins expressed in a given type of cell, tissue or organism under particular biological conditions at a given time. Shotgun (or bottom-up) proteomics is the most commonly used MS-based approach to study proteins by digesting proteins into peptides prior to MS analysis.
PEAKS Studio is a software platform with complete solutions for discovery and targeted proteomics, including protein identification and quantification, analysis of post-translational modifications (PTMs) and sequence variants (mutations), and peptide/protein de novo sequencing.
PEAKS Studio 12
Complete & Vendor Neutral Solution for Proteomics with DDA & DIA Support
PEAKS® Studio 12 offers a complete bottom-up proteomics solution with increased accuracy, sensitivity, and speed. Updated workflows for a variety of applications, such as in depth canonical and non-canonical peptide and protein identifications, make PEAKS® a unique solution. From DDA to DIA data support, PEAKS® Studio 12 provides a comprehensive solution to bring your research to new heights!
Contact us to add PEAKS Studio 12 to your lab!
Highlights
- Improved DeepNovo sequencing with FDR control.
- De novo assisted database search with a deep learning-boost to maximise peptide ID efficiency.
- DIA workflow (Spectral Library Search + directDB + de novo).
- Enhanced sensitivity and accuracy for direct database searches.
- Enhanced Deep learning-based technology for predicting spectra, retention time, and collision cross-section values.
- Hybrid DIA and targeted proteomics solutions.
- Post-translational modification (PTM) search with 500+ modification.
- Sequence variant and mutation search.
- DeepNovo-based peptidome workflow for immunopeptidomics: integrated canonical and non-canonical database searches, homology search, and DeepNovo for improved HLA peptidomics with FDR control.
- DIA Peptidome workflow.
- Label-free and Label-based: TMT (MS2, MS3) / iTRAQ, SILAC, 18D labelling, ICAT, User-Defined.
- Quantification results can be visualised using heat maps, correlation profiles, and extracted ion chromatograms (XICs).
- Detailed and easy-to-use graphical user interface (GUI) to view, filter and validate results.
- Spectral library viewer to assess the quality and validate library before use.
- Statistical calculations presented visually to assess quality of raw data and/or results.
- LC-MS/MS heat maps provide full visualisation of peptide features, MS/MS spectra acquisition, and identification position relative to mass over charge (m/z), retention time (RT), compensation voltages (CV), ion mobility (1/k0), and signal intensity.
- PEAKS IMS add-on module: Support for ion mobility mass spectrometry proteomics (timsTOF Pro, FAIMS).
- Algorithms fine tuned for each instrument and fragmentation type to ensure optimal accuracy and sensitivity.
- Comprehensive support of DDA and DIA for identification and quantification analyses.
Software Overview
Algorithms
PEAKS 12 uses the latest PEAKS algorithm for all analyses, including data loading/refinement, identification and quantification. New to PEAKS Studio 12 users have the option to use Deep learning-boosted ID workflows for DDA analysis to increase of identification rate over 10%.
Streamlined Workflow with Direct Database search for DIA
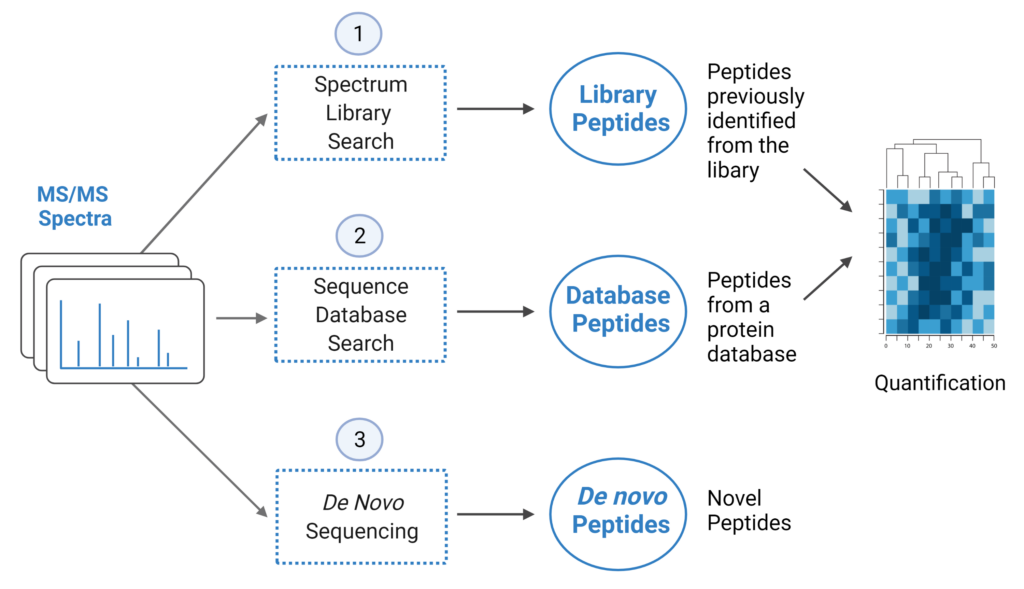
PEAKS Studio 12 offers a unique DIA workflow to maximise identification of peptides by integrating three methods: spectral library search, direct database search, and de novo sequencing, with up to 20% increase in identification and quantification.
- A library search is performed against a predefined spectral library. Peptide spectra without a library match can be directly searched against a protein database.
- A protein sequence database is directly searched with DIA data. Advanced machine learning algorithms allow improved accuracy and sensitivity of peptide identification. During this step of the pipeline, PEAKS 12 DIA workflow now supports the identification of any PTMs specified by the user. This will enable an increase in identification of modified peptides without requiring their entries in a spectral library.
- Unmatched spectra from the database search are de novo sequenced.
- Identified peptides from both the spectrum library search and protein sequence database search can be used in a quantification analysis.
Note: All 3 search methods, PEAKS Spectral Library Search, Direct Database Search, and de novo sequencing are optional, and the steps of the workflow can be conducted independently or in sequence.
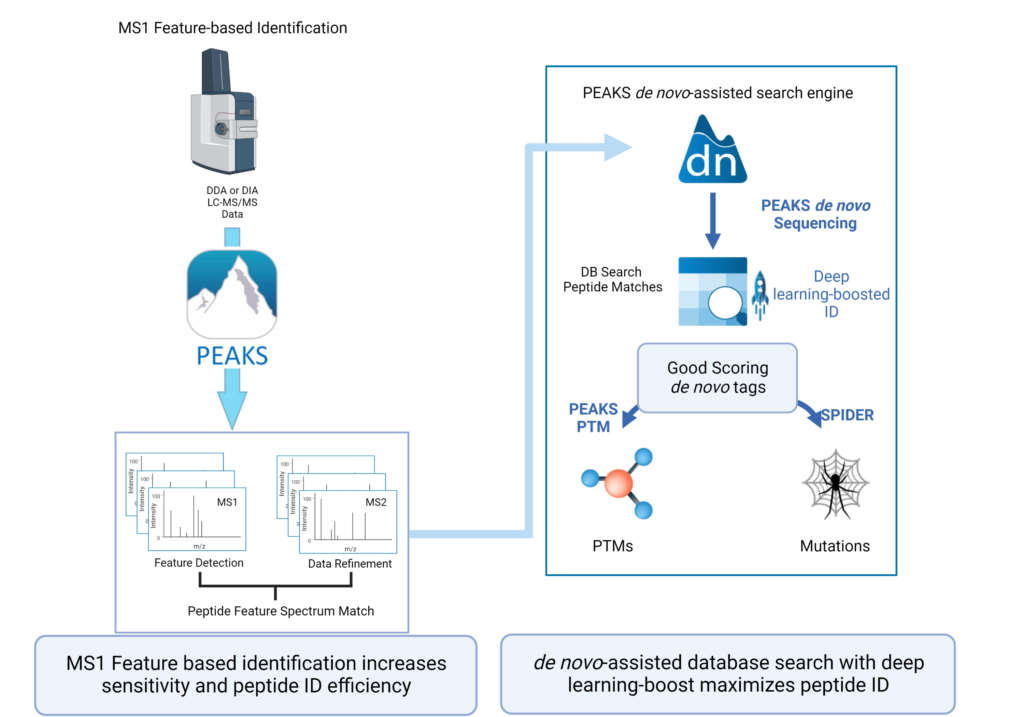
Feature-based identification workflow to increase sensitivity and maximise peptide identification efficiency.
- Designed for DDA technology to improve reproducibility.
- Integrate database search and de novo sequencing to extend in-depth analysis.
- Activate Deep learning-boost in PEAKS DDA workflows to maximise peptide ID efficiency.
Learn more about the advantages de novo sequencing brings to your research.
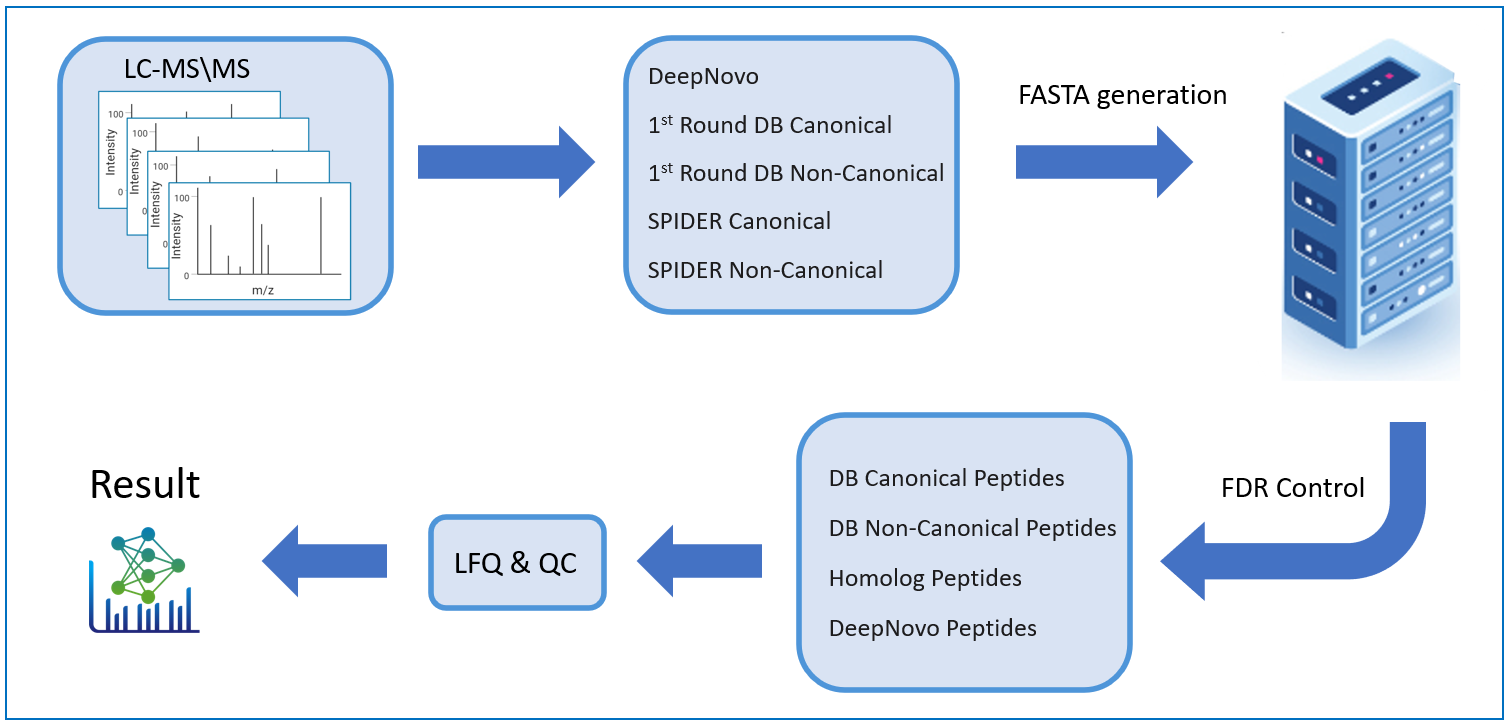
PEAKS DeepNovo Peptidome: Advanced solution Immunopeptidomics
This newly developed solution is a specialised workflow for peptidomics data that combines database searching, de novo sequencing, and identification of mutated peptides. Learn more
Ion mobility enabled quantification to minimise missing-values and enhance accuracy
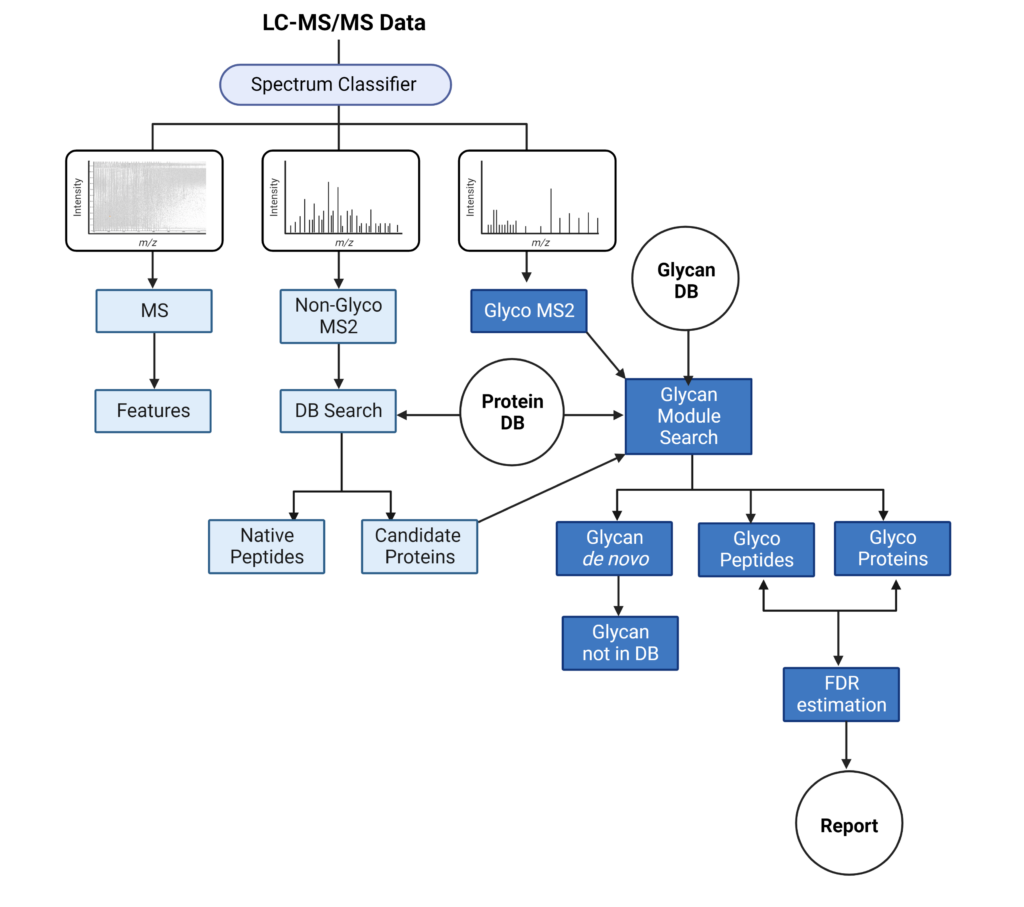
PEAKS Glycan search enables scientists to determine glycan site localisation and glycan structures
PEAKS Glycan search provides a highly sensitive and accurate glycoproteomics software solution to advance our understanding of the glycoproteome. Learn more
- Comprehensive understanding of glycosylation and glycoproteins
- Advanced glycoproteomic for deep glycan profiling
- Innovative software tool for glycan and glycopeptide quantification
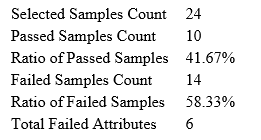
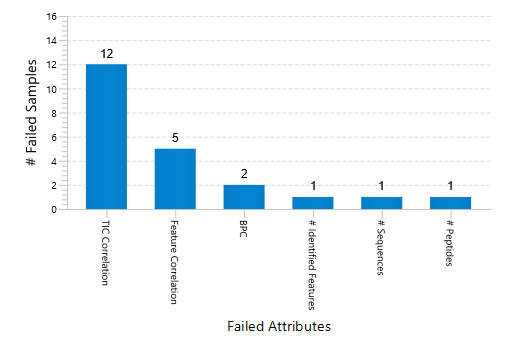
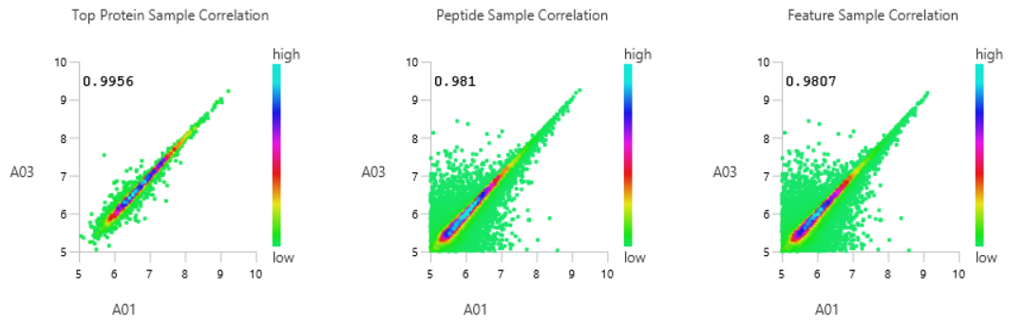
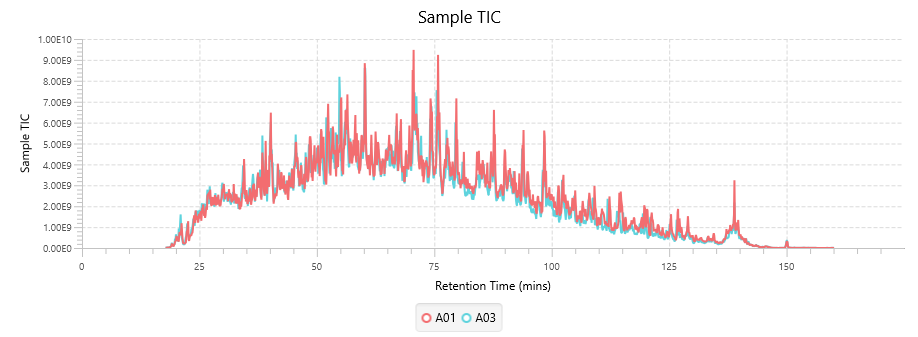
Quality Control (QC) Function for in-depth analysis from raw data to results
With the new Quality Control (QC) analysis in PEAKS® Studio 12 users can assess statistical information of the raw data and/or results and gain beneficial insight into the attributes of the LC-MS acquisition. This automated tool is designed for both, DDA and DIA data and will supply the elements to determine the quality of the data and evaluates the setup of the experiment.
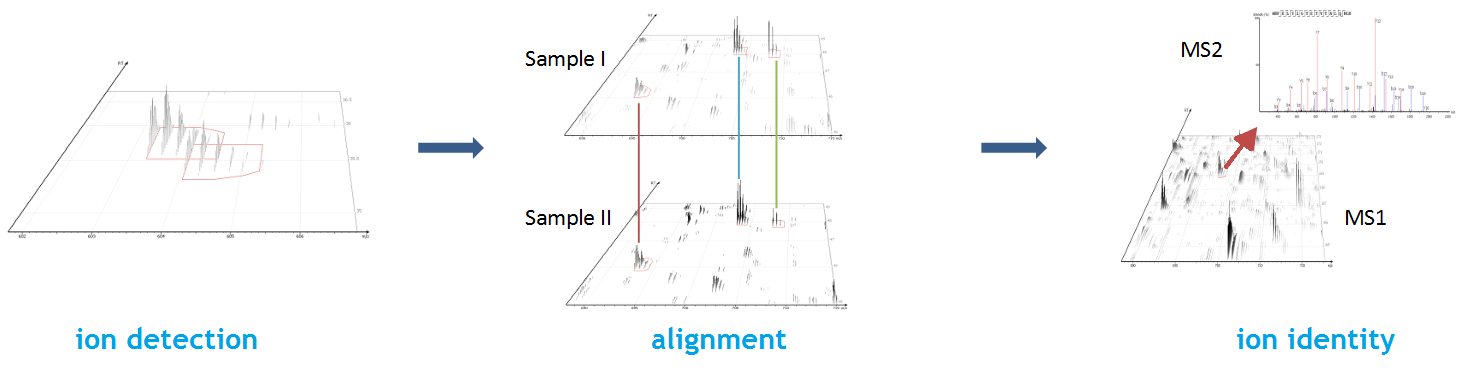
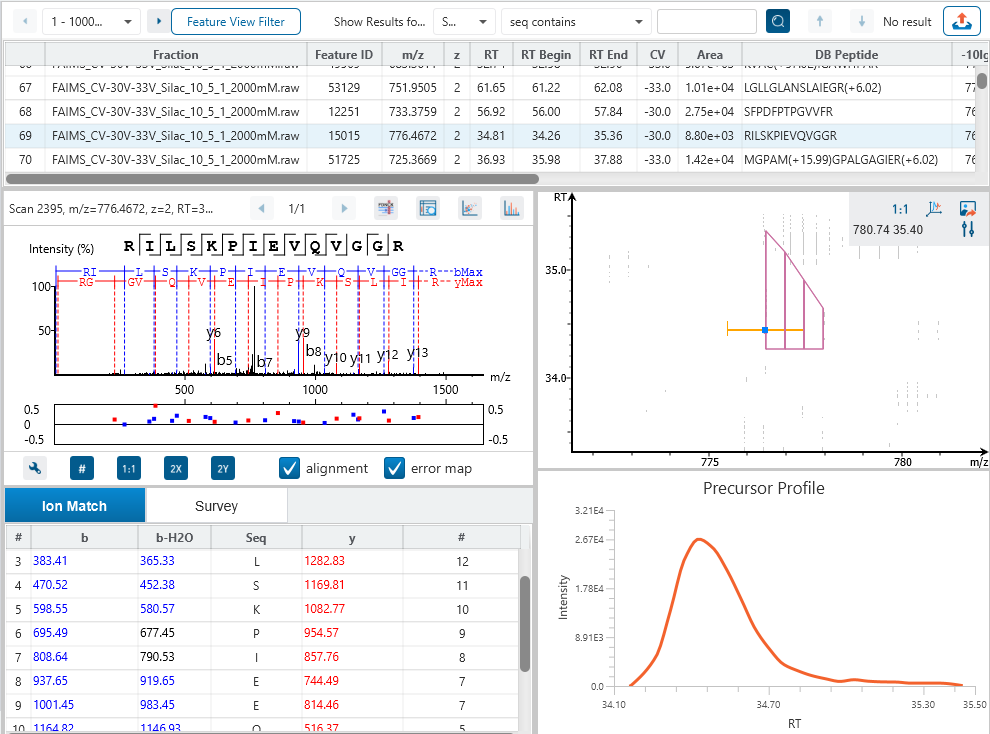
The Peptide Feature View
From data refinement to identification and quantification, the PEAKS 12 series is peptide feature-based. A peptide feature includes a series of corresponding m/z values, a retention time range, the intensities formed by different molecular isotypes, and the ion mobility as the fourth dimension information.
In PEAKS identification results, users will find a “Feature” tab which presents all details of every peptide feature in the raw data, visually and tabularly. In the Peptide Feature table, PEAKS conveniently summarises feature details detected from LC-MS or LC-IMS/MS, including the identified sequence as determined by database search, DDA, or de novo sequencing. Each feature is also linked to their corresponding protein view, spectrum view and LC-MS or LC-IMS/MS view for further inspection.
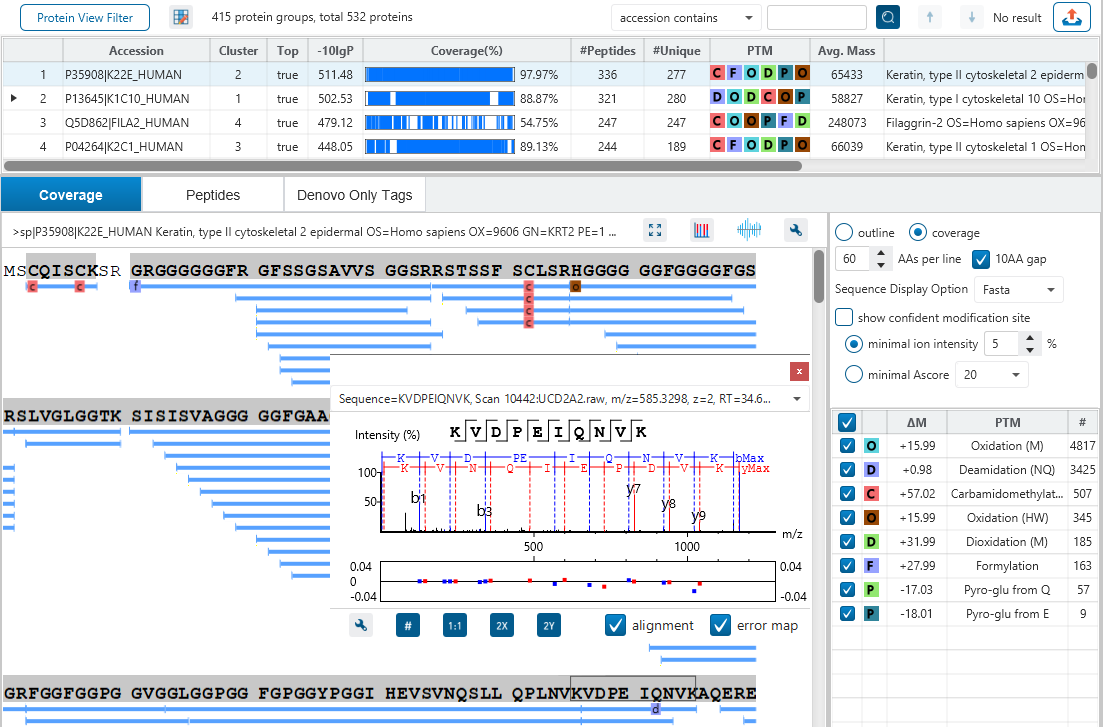
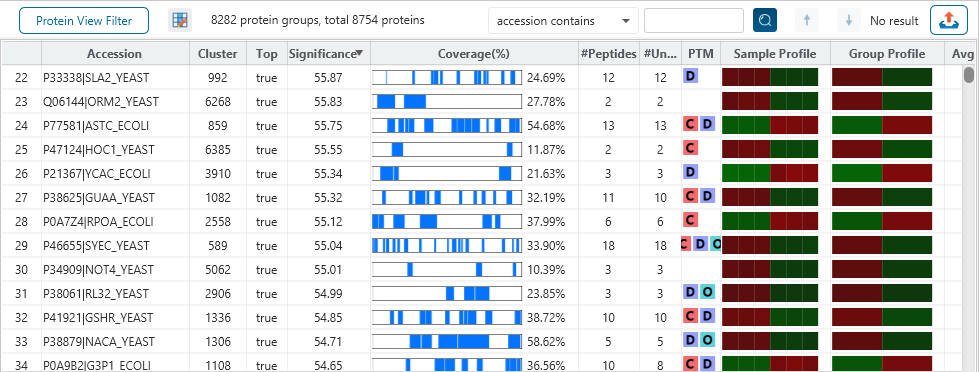
The Protein View
The Protein View presents protein profiling across complex biological samples in both DDA and DIA workflows. For each protein, the Sequence Coverage View displays a peptide map with spectrum annotation for validation. With PEAKS traditional de novo-assisted database search, users easily view identified peptide sequence in blue, while a grey bar indicates a de novo only tag match.
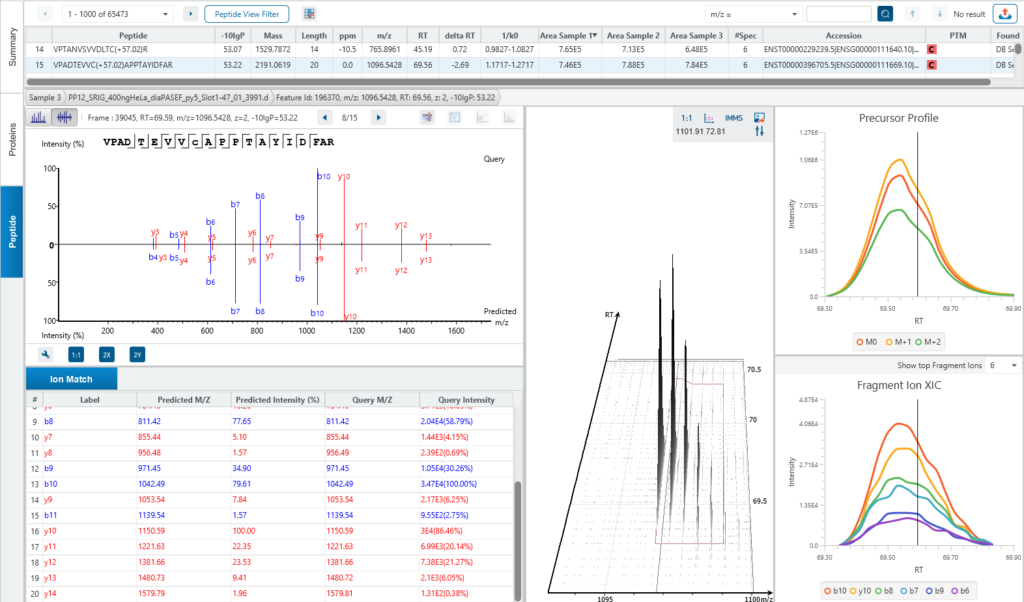
The Peptide View
The Peptide View provides a list of identified peptides with the abundance from MS1. For each modified peptide, the confidence of modification site (Ascore) is associated.
The Quantification View
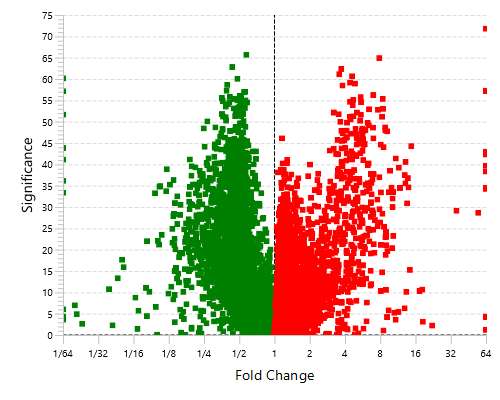
With the add-on module of PEAKS Q, PEAKS Studio also determines relative protein abundance changes across a set of samples simultaneously and without the requirement for prior knowledge of the proteins involved.
Highly differentially expressed proteins between two groups are identified by statistical analysis tool (fold change >2, FDR <0.01) and displayed in a heatmap format.
Licence Information
The PEAKS Studio licence can be scaled to address your lab’s requirements.
- Desktop – 24 threads, capable of processing across up to 24 cores
- Workstation – 48 threads, up to and across 48 cores
Configuration
PEAKS Studio 12 is recommended to be installed on 64-bit Windows operating systems with Windows 10 or later.
Recommended Configurations:
Desktop License: 30+ threads processors and 64 GB+ of RAM with compatible GPU (described below)
Workstation License: 60+ threads processors and 128 GB+ of RAM with compatible GPU (described below)
e.g. Intel Core i7/i9/Xeon or AMD Ryzen 7/9/Threadripper processors
For running DIA Database Search, it is required that the machine is equipped with a GPU having an NVIDIA CUDA compute capability >= 5 with at least 8GB of dedicated memory.
For running DeepNovo, it is required that the machine is equipped with a GPU having an NVIDIA CUDA compute capability >= 8 with at least 8GB of dedicated memory.
GPU compute capability can be found on NVIDIA’s website: https://developer.nvidia.com/cuda-gpus
Furthermore, the GPU needs to be updated to CUDA version 12.3 or later. Please find the CUDA Toolkit installer on NVIDIA’s website and also install the packaged NVIDIA driver.
References & Resources
References
- Tran NH, Qiao R, Xin L, Chen X, Liu C, Zhang X, Shan B, Ghodsi A, Li M, Deep learning enables de novo peptide sequencing from data-independent-acquisition mass spectrometry, Nat. Methods, 16, 63–66 (2019). https://doi.org/10.1038/s41592-018-0260-3
- Tran NH, Zhang X, Xin L, Shan B, Li M. De novo peptide sequencing by deep learning. Proc. Natl. Acad. Sci. U.S.A. 114, 8247-8252 (2017). https://doi.org/10.1073/pnas.1705691114