BSI Lab Services

Moving your science forward
At Bioinformatics Solutions Inc., we focus on proteomic analysis using mass spectrometry and bioinformatics to provide comprehensive identification and characterization of proteins and glycans.
Proteomics and glycomics are the study of the repertoire of proteins and glycans expressed in a cell, tissue, or organism under a particular biological condition at a given time. The study of these biomolecules have a wide range of applications in basic and applied science including cellular and systems biology, medicine, drug discovery, forensics, personal care products, food sciences and so much more. Importantly, the application of proteomics and glycomics provides major opportunities to elucidate disease mechanisms and identify new diagnostic biomarkers and novel therapeutic targets.
Request Pricing / Schedule Consultation
What do we provide?
Bioinformatics Solutions Inc. offers cost-effective services for mass spectrometry (MS) based proteomics with fast turnaround times (1-3 weeks). As a Software company and a Contract Research Organisation, we can perform all sample preparation and complex mass spectrometry experiments in-house using our state-of-the-art equipment and facility. The data is then analysed using our PEAKS software. Data analysis is included with every experiment. Additionally, our services are individually customised and tailored to your specific needs.
Why use our service?
With over 20 years experience in the proteomics research community, we understand the importance and value of proteomics and glycomics research to advance our understanding of the complexity underlying biological systems. We provide fast turnaround times, complete confidentiality, and guaranteed results through cost-effective and efficient mass spectrometry solutions for a wide range of industries.
Contact us for a personal consultations to develop your project, and perform all experiments in a professional, accurate, and timely manner.
BSI Lab Services
Protein Identification and PTM Analysis
Service Description:
We process in-solution or in-gel samples and offer customised digestion with 5 different enzyme options available to enhance protein coverage. Liquid chromatography tandem mass spectrometry (LC-MS/MS) is used to generate raw data. Our latest PEAKS Studio 11 software is used for both de novo sequencing and database searching to provide sensitive and accurate peptide/protein identification results. A targeted search for any post-translational modifications (PTMs) in PEAKS Studio 11 can be included as an optional add-on service. Our user-friendly PEAKS Studio Viewer (included) provides an easy way to further investigate your data and extract images of the results.
Workflow:
In-solution/in-gel digestion → LC-MS/MS → Data analysis
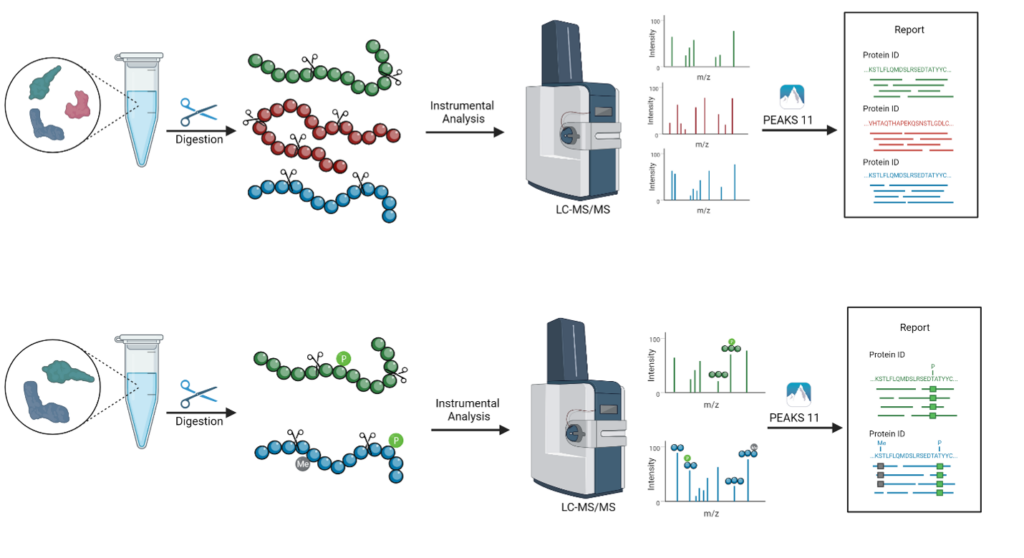
What we need:
- 20μg of sample provided in-solution or as gel bands
- Samples must be free of detergents and polymers (Triton X-100, Tween-20, PEG, etc.)
- Buffer information for in-solution samples
What we deliver:
- PEAKS Studio project file containing database search results and PTM analysis
- PEAKS Studio Viewer
- Raw data files
Antibody-Drug Conjugate Analysis
Service Description:
When producing or developing antibody-drug conjugates, defining the drug-to-antibody ratio and conjugate site-specificity are critical quality control steps. In addition to retrieving an unknown antibody sequence, this service includes intact mass deconvolution to analyze drug-to-antibody ratios and PTM analysis to identify the conjugation site(s).
Workflow:
In-solution digestion → LC-MS/MS → Antibody sequencing and PTM analysis
Deglycosylation → LC-MS → Intact mass deconvolution
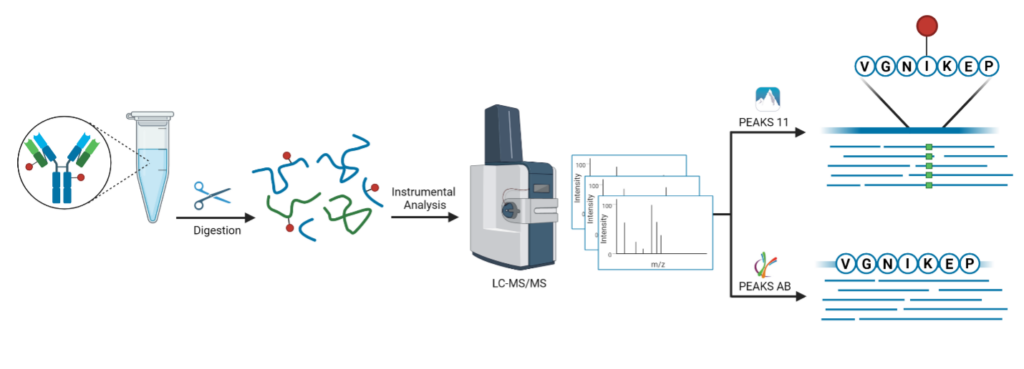
What we need:
- 100μg of sample provided in-solution
- Samples must be free of detergents and polymers (Triton X-100, Tween-20, PEG, etc.)
- Buffer information
- Mass of the conjugate (for peptide-based conjugates, we require mass of conjugate after protease digestion)
What we deliver:
- PEAKS ADC Report
- Antibody sequence, intact mass analysis, peptide mapping, I/L differentiation, MS2 spectra covering variable region, MS2 spectra showing conjugation site
- PEAKS Studio Viewer
- Raw data files
Intact Mass Analysis
Service Description:
In-solution samples are analysed by LS-MS to measure purified proteins at the intact mass level. The samples can be processed under different treatment conditions (reduction, alkylation, deglycosylation, etc.) and theoretical protein masses are matched to peaks in deconvoluted spectra.
Workflow:
Sample preparation → LC-MS → Intact mass deconvolution → protein matching and peak annotation
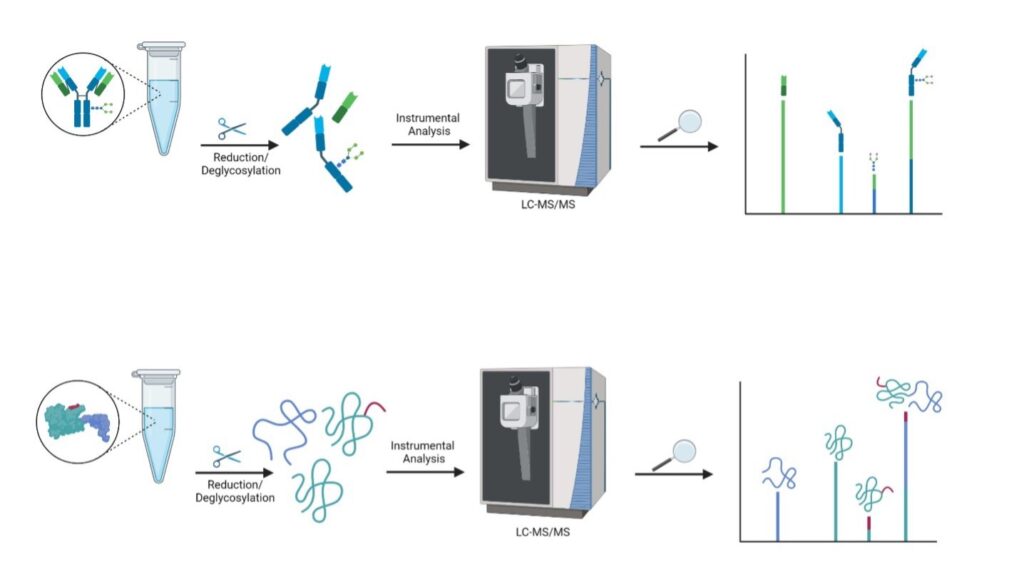
What we need:
- 20μg of sample provided in-solution
- Samples must be free of detergents and polymers (Triton X-100, Tween-20, PEG, etc.)
- Buffer information
What we deliver:
- PEAKS Intact Report
- Chromatogram, original spectrum, deconvoluted spectrum
- Proteins matching to intact mass peaks
- Raw data files
Glycan Profiling and Glycopeptide Analysis
Service Description:
Protein glycosylation is one of the most common PTMs and directly affects the structure and function of a protein. Identifying glycans and determining protein glycosylation sites is crucial information for characterising a protein and developing therapeutics. In this service we identify N- and O-linked glycans along with protein glycosylation sites using our PEAKS GlycanFinder. We process in-solution or in-gel samples and offer glycopeptide enrichment for more complex samples.
See how PEAKS GlycanFinder can move your science forward and learn more details from our application notes that focus on samples runs in our lab.
- N-glycan and O-glycan Profiling of Fetuin by single LC-MS/MS run
- Characterisation of N-linked glycosylation patterns of IgG antibodies in PEAKS GlycanFinder
- Identifying different glycan profiles of CD33 expressed in HEK293 and CHO cells
Workflow:
In-solution or in-gel digestion → LC-MS/MS → Glycan profiling/identification → Quantification (optional)
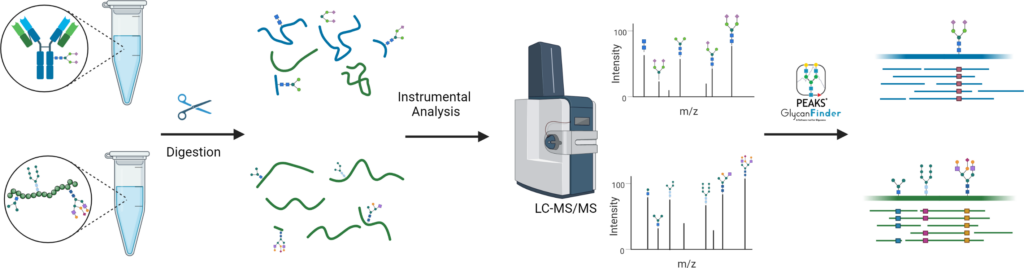
What we need:
- >10μg of sample provided in-solution or in-gel
- Samples must be free of detergents and polymers (Triton X-100, Tween-20, PEG, etc.)
- Buffer information
- Target protein information (species or cell line)
Type of analysis:
- Glycan Identification for purified proteins with intact mass validation
- Intact sample preparation and mass deconvolution
- Tryptic digestion and LC-MS/MS on timsTOF Pro (3 replicates per sample)
- Match to top 5 most abundant glycoforms to intact masses
- Glycan Report I
- Glycoproteomics – glycan identification of complex samples
- Tryptic digestion, glycopeptide enrichment, and LC-MS/MS on timsTOF Pro (3 replicates per sample)
- Glycan Report II
What we deliver:
- PEAKS Glycan Report
- Based on the type of analysis we will either provide:
- Glycan Report I – Glycoprotein sequence, intact mass results with peak annotation to top glycoforms, peptide mapping of N-linked and/or O-linked glycopeptides, list of all identified glycans and their glycosylation sites, MS2 spectra of top supporting glycopeptides
- Glycan Report II – Glycoprotein sequences, peptide mapping of N-linked and/or O-linked glycopeptides to top 5 identified glycoproteins, list of all identified glycans and glycoproteins (provided as .csv file), MS2 spectra of top supporting glycopeptides for the top 5 glycoproteins identified
- Based on the type of analysis we will either provide:
- Raw data files

